Folia Microbiol. 33, 462-465 (1988)
of Trichomonas vaginalis
J. FLEGRa, J. ČERKASOVb and J. ŠTOKROVÁa
aInstitute of Molecular Genetics, Czechoslovac Academy of Sciences, 166 37 Prague 6, Faculty of Science, Charles University, 128 44 Prague 2
Received February 13, 1988
ABSTRACT. The existence of six dsRNA segments of Trichomonas vaginalis virus was confirmed and the molar mass and relative abundance of these segments ware determined by agarose gel electrophoresis with reovirus dsRNA serving as a standard. The M's were 3.5, 3.4, 3.2, 2.5, 1.4 and 0.34 Mg/mol for the two strains studied, the relative abundances, however, were 1.0, 1.4, 3.0, 0.3, 2.7, 4.2 and 1.0, 0.6, 1.7, 0.5, 3.4 1.0 for these strains, respectively. Cell homogenate fractionation showed that all dsRNA segments were associated with viral particles. The data appeared to support the hypothesis of a relationship between viruses of the protozoan T. vaginalis and of the yeast Saccharomyces cerevisiae.
Trichomonas vaginalis, a sexually transmitted protozoan parasite of the human urogenital tract, has been found to harbor a dsRNA virus with an icosahedral capsid 33 nm in diameter (Wang and Wang 1986; Flegr et ad. 1987a). The virus is rather common, its dsRNA being detectable in 45-95 oo of T. vaginalis strains (Flegr et al.1987a; Wang and Wang 198). The presence of the virus appears to correlate with the mode of expression of a dominant T. vaginalis antigen (Wang et al. 1987). Trichomonads of virus-harbouring strains express this antigen on their surface, trichomonads of virus-free strains contain it in their cytoplasm only. Absence of this antigen on the cell surface has been claimed to be a marker of certain virulence attributes, namely cytoadherence and cytotoxicity (Alderete et al. 1986; Alderete and Garza 1985). No differences, however, were found between dsRNA negative and positive strains if the virulence was judged on the basis of mouse assays and clinical and histopathological manifestations on female patients (Flegr et al. 1987cc).
In the size and shape of the capsid, in the molar mass of its most abundant protein, as well as in the dsRNA nature of its genome, the T. vaginalis virus resembles the killer viruses of Saccharomyces cerevisiae (Tipper and Bostian 1984). To estimate the extent of this similarity, reliable data on the dsRNA molecules of the T. vaginadis virus are required. Wang and Wang ( 1985,1986) reported the presence of a single species of dsRNA in T. vaginalis. In contrast, our previous results demonstrated the presence of multiple populations of dsRNA (Flegr et al. 1987b).
In this paper ve confirm that several dsRNA species can be extracted from viral particles isolated from T. vaginali,s and report the NI's and relative abundance of the six dsRNA species.
MATERIAL AND METHODS
Organisms. Axenic Trichomonas vaginali strains Al (Meingassner and Thurner 1979) and TV 10-02 MR (Čerkasovová et al. 1987) were used as sources of dsRNA. Trichomonads were stored as cryostabilates ( % dimethyl sulfoxide, liquid nitrogen) and cultivated without antibiotics in tryptose-yeast extract-maltose (TYM) medium (Diamond 1957) supplemented with 10% horse serum.
Isolation of viruses. Trichomonads (cell concentration 40nL in 0.8 oo NaCl) were disintegrated in a Potter-Elvehjem homogenizer, diluted 1 : 20 with 0.8 % NaCl and centrifuged (20 min.,15000 g). The pellet was resuspended in the original volume of 0.8 % NaCl and centrifuged as before. The supernatants were pooled and viral particles (VP) were obtained from this pool by ammonium sulfate fractionation (solid ammonium sulfate, saturation 20, 30, 40, 50 and 90 %, each step lasting 2 h at 0 oC). The highest quantity of VP precipitated between 30 and 40 oo saturation (Plate 1 ).
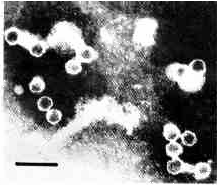
Plate 1
Viral particles isolated by ammonium sulfate precipitation. Negative staining with phosphothungstic acid (Frank et al. 1978). Bar represents 100 nm.
Isolation of nucleic acids. Nucleic acids were extracted with a chloroform3-methyl-1-butanol mixture (24 :1 ) in the presence of guanidine hydrochloride (4 mol/L) and precipitated with a 0.7 volume of 2-propanol (Flegr 1987). ,
Electrophoresis. Nucleic acids were electrophoresed in 1% agarose gel in Tris-borate buffer and detected with ethidium bromide (Maniatis et al.1983). Alternatively, nucleic acids were stained before electrophoresis with 4',6- diamidino-2-phenylindol. (DAPI) or with a mixture of DAPI and ethidium bromide (3 and 15 ppm). Densitograms were read from negatives with a Magiscan 1 analyzer (Joyce-Loebl). The M's of dsRNAs, supposed to be proportional to the inverse values of their electrophoretic mobility (Edmanndson and Gray 1984), were estimated by electrophoresis with a dsRNA standard (reovirus type III, strain Dearing). The relative abundance of the segments were calculated from their M's and from the intensity of fluorescence of the electrophoretic zones.
RESULTS
The extracted nucleic acids contained DNA (zone a), ssRNA (zone b) and six other distinct populations of nucleic acids (zones I-VI) (Plate 2, 3). Zones I-VI were susceptible to pancreatic ribonuclease (RNAse A; EC 3.1.27.) (Plate 4) but resistant to ribonuclease T1 (RNAse Tl; EC 3.1.27.3) (Plate 5) and deoxyribonuclease I (DNAse I; EC 3.1.21.1) (Plate 6): When the nucleic acids were stained with DAPI, zones I-VI as well as the DNA zone emitted bluish fluorescence. When a mixture of DAPI and ethidium bromide was used, zones I-VI and the DNA zone emitted a red and a bluish fluorescence, respectively. All these results indicated that zones I-VI consisted of dsRNA.
Plate 2. Nucleic acids from two strains of T. vaginalis. a- DNA, b - ssRNA, I-VI dsRNA. Lanes 1,2 - strain TV 10-02 MR5, 4,5 - strain A1, 3 - DNA of phage lambda digested with HindIII.
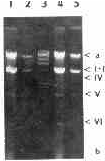
Plate 3. The same electrophoretic gel as on Plate 1, after prolonged electrophoresis.

Plate 4. Nucleic acids from T. vaginalis strain A1 digested with different concentrations of RNAse A before electrophoresis. From left to right, 0, 0.13, 2, and 32 mg of enzyme per mL of medium incubation buffer (Maniatis et al. 1983).

Plate 5. Nucleic acids from T. vaginalis strain A1 digested with different concentrations of RNAse T1 before electrophoresis. From left to right, 0, 2, 32 and 512 mg of enzyme per uL of medium incubation buffer (Maniatis et al. 1983).

Plate 6. Nucleic acids from T. vaginalis strain A1 digested with different concentrations of DNAse I before electrophoresis. From left to right, 3.2, 12.8, 51 and 205 ug of enzyme per uL of medium incubation buffer (Maniatis et al. 1983).

The molar mass and relative abundance of dsRNA from strain Al were 3.5, 3.4, 3.2, 2.5, 1.4, 0. 34 Mg/mol and 1.0, 1.4, 3.0, 0. 3, 2.7, 4.2, respectively. The M's of the dsRNA segments from strain TV 10-02 MR were the same, but their relative abundance were 1.0, 0.6, 1.7, 0.5, 3.4 and 1.0.
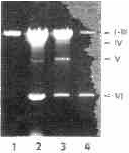
Extracts of nucleic acids from purified VP differed from those obtained from whole protozoans by the absence of DNA and by a lesser amount of ssRNA. The presence of all six dsRNA species, however, was clearly apparent. A maximum of dsRNA was detected in preparations of VP precipitated with 40 o/a saturation of ammonium sulfate; the relative abundances of particular dsRNA species, however, varied between the different fractions of VP obtained by ammonium sulfate fractionation (Plate 7).
Plate 7. Nucleic acids isolated from viral particles precipitated with 30, 40. 50 and 60% saturated ammonium sulphate (lanes 1, 2, 3, and 4, respectively).
DISCUSSION
In the present study, the existence of six dsRNA species in the viral particle fraction from T. vaginalis was confirmed. The M's and the relative abundances of the particular species were estimated and an interstrain variability of the latter was demonstrated.
Wang and Wang (1985, 1986) report the presence of a single dsRNA species only. We suppose that differences in experimental methods rather than interstrain variability are responsible for this discrepancy, because: 1. No variability in the number of dsRNA species has been reported, in spite of the fact that large collections of T. vaginalis strains harbouring dsRNA from different parts of the world have been studied (Flegr et al. 1987a: Wang and Wang 198; Wang et al. 1987). 2. Wang and Wang used guanidine isocyanate instead of guanidine hydrochloride for nucleic acid isolation. Both methods were compared and it was found that the former gives only scant yields of dsRNA so the less abundant species of dsRNA can easily escape detection (Flegr 1987). Furthermore, the electromobilities of the three largest dsRNA segments were very similar and long agarose gels or a polyacrylamide gel had to be used for their separation (compare Plate 2 with Plate 3). The different abundance of particular dsRNA segments and variation in the relative amounts of particular dsRNAs in different VP fractions or in different trichomonad strain suggested that at, least some segments were packed separately, in distinct VP.
The existence of several dsRNA species of different 1M's and abundance, the range of their M's, as well as the existence of interstrain variability in the abundance, resembled the situation with killer viruses of S. cerevisiae (Tipper and Bostian 1984). However, direct evidence confirming the hypothesis of a relationship between the trichomonad and the yeast viruses suggested by Wang and Wang ( 1986) is still missing.
The authors express their thank to Dr. J. Kulda (Prague) for providing the trichomonad cultures used in the experiments, to Dr. J.G. Maingassner (Viena) for the T. vaginalis strain Al and to Dr. M. Rosembergová (Bratislava) for the reoviral dsRNA.
REFERENCES
ALDERETE J.F., KASMALA L., METCALFE E., GARZA G.E.: Phenotypic variation and diversity among Trichomonas vaginalis and correlation of phenotype with trichomonal virulence determinants. Infect.Immun. 3, 285-93 (1986).
ALDERETE J.F., GARZA G.E.: Specific nature of Trichomonas vaginalis parasitism of host cell surfaces. Infect. Immun. 50, 701- 708 ( 1985).
ČERKASOVOVA A., NOVAK J., CERKASOV J., KULDA J., TACHEZY J.: Metabolic properties of Trichomonas vaginalis resistant to metronidazole under anaerobic conditions, in Trichomonads and Trichomoniasis, Part 2 (J. Kulda, J. Carkasov, eds.). Acta Univ.Carolinae Biol. 30, 505-512 (1986).
DIAMOND L.S.: The establishment of various trichomonads of animals and man in axenic cultures. J.Paasitol. 43, 488-490 (1957).
FLEGR J., CERKASOY J., KULDA J., TACHEZY J., STOKROVA J.: The dSRNA Of Trichomonas vaginalis is associated with virus-like particles and does not correlate with metronidazole resistance. .Folia Microbiol. 32, 345-348 (1987a).
FLEGR J., CERKASOV J., CERKASOVOVA A., KULDA J., STOKROVA J.: Double stranded RNA in Trichomonas vaginalis, in Trichomonads and Trichomoniasis, Part 1 (J. Kulda, J. Cerkasov, eds.). Acta Univ. Carolinae Biol. 30, 281-286 (1987b).
FLEGR J.: A rapid method for isolation of double stranded RNA. Prepar.Biochem. 17, 433-433 (1987).
FRANK H., SCHWARZ H., GRAF T., SCHAFER W.: Properties of mouse leukemia viruses. XV. Electron microscopic studies on the organisation of Friend leukemia virus and other mammalian C-type viruses. Z.Naturforsch. 33c, 124-128 (1970).
EDMArrrrDsOrr S.P., GRAY D.M.: Analysis of electrophoretic properties of double stranded DNA and RNA in agarose gels at a finite voltage gradient. Biopolymers 23, 2725-3742 (1984).
MANIATIS T., FRITSCH E.F., SAMBROOK J.: A Laboratory Manual, Molecular Cloning, Cold Spring Harbor Laboratory: Cold Spring Harbor, New York 1983.
MEINGASSNER J.G., THURNER J.: Strain of Trichomonas vaginalis resistant to metronidazole and other 5-nitroimidazoles. Antimicrob.Agents Chemother. 15, 254-257 (1979).
TIPPER D.J., BOSTIAN K.A.: Double stranded ribonucleic acid killer systems in yeasts. Microbiol. Rev. 48, 125-156 (1984).
WANG A.L., WANG C.C.: A linear double stranded RNA in Trichomonas vaginalis. J.Biol.Chem. 260, 3697-3702 (1985).
WANG A.L., WANG C.C.: The double stranded RNA in Trichomonas vaginalis may originate from virus-like particles. Proc.Nat.Acad.Sci. USA 83, 7956-7961 (1986).
WANG A.I.., WANG C.C., ALDERETE J.F.: Trichomonas vaginalis phenotypic variation occurs only among trichomonads infected with the double-stranded RNA virus. J.Ep.Med.166,142-150 ( 1987).